Saravanan Palani

Saravanan Palani
Assistant Professor
Ph.D: German Cancer Research Center (DKFZ), Germany
Post-doctoral research:
University of Warwick, United Kingdom
Year of Joining: 2020
Phone: +91 80 22932306
Email: spalani@iisc.ac.in
The focus of our lab is to decipher how cytoskeletal forces are generated through actomyosin ring (AM-r) machinery during cell division.
Highlights
- Studying how the components of the AM-r assemble at the divisional apparatus at the correct time during cell cycle.
- Deciphering how the AM-r is anchored to the membrane.
- Understanding the novel mechanisms of how AM-r generates a force in the presence or absence of motor protein Myosin-II to divide the cells.
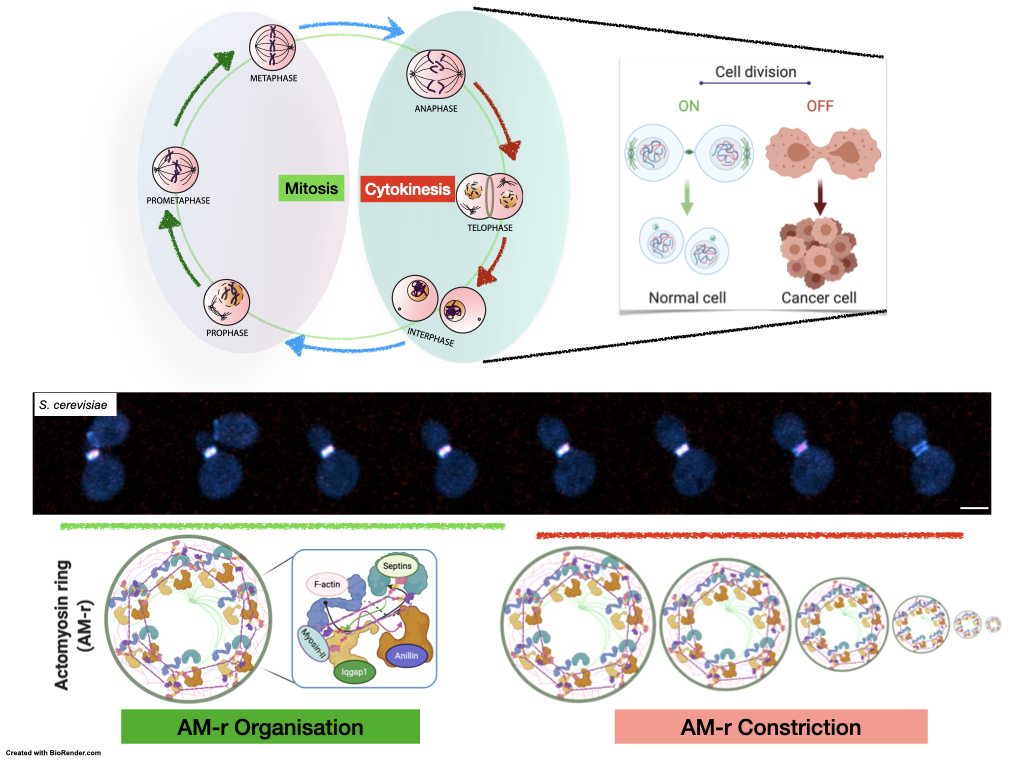
Deciphering the mechanism of ActoMyosin-ring (AM-r) assembly and constriction. Schematic representation depicts the cell cycle regulation of the cytokinesis; time-lapse microscopy images show sequential events of the AM-r assembly and constriction in budding yeast (S. cerevisiae).
Mechanical forces are vital in driving a variety of cellular processes such as cell division, migration, wound healing and contraction of muscle cells. One of the most crucial biological nano-machines powering these events is the actomyosin machinery network(Szent-Gyorgyi A., 1942 and 1943; Huxley., 1954; Meitinger and Palani., 2016). This machinery is mainly comprised of actin filaments (F-actin), the molecular motor Myosin-II, Tropomyosin and actin cross-linkers. The interplay between these proteins, primarily the relative sliding between myosin-II and actin filaments, leads to tension generation in the actomyosin network. Though the actomyosin machinery is conserved across all eukaryotes - from yeast to humans - a comprehensive understanding of the underlying mechanisms that drive this machinery is still sought after. For example, the core features and properties shared by circular actomyosin ring assemblies are not fully understood. Likewise, the mechanisms underlying actomyosin ring functions in different cellular contexts such as wound repair, phagocytosis, T-cell synapse, and cytokinesis remain elusive.
Regulation and Dynamics of Cytokinetic Actomyosin Machinery: The main bottleneck to translate the functioning of actomyosin machinery in multiple biological settings is a lack of tools to follow the dynamics of the process at the cellular and molecular level. More broadly, our research will address current technical limitations in understanding the AM-r dynamics and regulation. To overcome these limitations, we will employ cutting-edge interdisciplinary approaches combining cell biology, synthetic biology (using non-canonical amino acids and click chemistry), fluorescence microscopy, biochemistry, molecular genetics and novel in vitro assays based on reconstituted lipid bilayer to address the open questions underlying actomyosin ring functions in various cellular contexts – yeast models (Saccharomyces cerevisiae and Schizosaccharomyces pombe) and mammalian cells.
Mechanisms of Actin Ring-Mediated Processes: In the long term, fundamental knowledge gained from deciphering the key aspects of actomyosin machinery will be used to delineate the role of actin ring-mediated processes that operate during wound repair in invertebrate model systems. This, in turn, will better our understanding of human diseases that are tightly linked with actomyosin-dependent regulation, such as cancer, neuronal disorders, and vascular disease. Overall, these studies will advance our knowledge of the actomyosin contractome network and its role in different cellular processes from single cell wound repair to embryonic development.
- DBT-Wellcome trust India Alliance Intermediate fellowship (2022-2026)
- DST-SERB-Startup Research Grant (2022-2024)
- CCAMP-InDx grant by The Rockefeller foundation (2020-2023)
Updating Shortly..
- Akhuli D, Dhar A, Viji AS, Bhojappa B, Palani S “ALIBY: ALFA Nanobody-Based Toolkit for Imaging and Biochemistry in Yeast” mSphere, 2022 DOI: https://doi.org/10.1128/msphere.00333-22 (MBoC Highlighted Preprint)
- Hatano T, Lim TC, Chaumartin IB, Dhar A, Gu Y, Wu TM, Scott W, Adishesha S, Chapa-y-Lazo B, Springall L, Sivashanmugam L, Mishima M, Martin SG, Oliferenko S, Palani S**, Balasubramanian MK** “mNG-tagged fusion proteins and nanobodies to visualize tropomyosins in yeast and mammalian cells” Journal of Cell Science, 2022 (**co-corresponding authors)
- Palani S, Ghosh S, E Ivorra-Molla, S Clarke, A Suchenko, Balasubramanian MK, Koster D “Calponin-Homology Domain mediated bending of membrane associated actin filaments” eLife, 2021. DOI: https://doi.org/10.7554/eLife.61078 (Recommended by Faculty opinions)
- Palani S**, Koster D, Hatano T, Kamnev A, Kanamaru T, Brooker H, Hernandez-Fernaud JR, Jones A, Millar JB, Mulvihill DP, Balasubramanian MK** “Phospho-regulation of tropomyosin is crucial for actin cable turnover and division site placement” Journal of Cell Biology, 2019. DOI: http://dx.doi.org/10.1083/jcb.201809089 (**co-corresponding authors)
- Palani S**, Chew TG, Ramanujam S, Kamnev A, Harne S, Chapa-Y-Lazo B, Hogg R, Sevugan M, Mishra M, Gayathri P, Balasubramanian MK** “Motor activity dependent and independent functions of myosin II contribute to actomyosin ring assembly and contraction in Schizosaccharomyces pombe”, Current Biology, 2017(b). DOI: http://dx.doi.org/10.1016/j.cub. 2017.01.028 (**co-corresponding authors)
- Meitinger F**, Palani S**. “Actomyosin ring driven cytokinesis in budding yeast”, Semin Cell Dev Biol, 2016. doi: https://doi.org/10.1016/j.semcdb.2016.01.043 (**co-corresponding author)
Ph.D. Students : 4
Mr. Anubhav Dhar (Aug 2020)
Ms. Bindu B (Aug 2020)
Ms. Freya Cardozo (Aug2022)
Ms. Jayanthi Kumari (Aug2022)
Openings:
We are always looking for promising and enthusiastic postdoc candidates with a background in Cell biology or Biochemistry, keen to work on interdisciplinary projects linked to the core research interests of the lab, in collaboration with other groups at IISc, NCBS and beyond
Please also take a look at other post-doctoral funding opportunities: DBT-Research Associate fellowship (DBT-RA) programme and the DBT-Wellcome India Alliance Early career fellowship.
PhDs: Apply through IISc admissions portal to the research programmes.
Our research group is currently looking for project assistants, JRFs and PhDs.
Interested candidates (Project assistants, JRFs and Post-docs) can please email a brief motivation letter outlining relevant expertise and scientific interests, theirCV, and contact information for three references to spalani@iisc.ac.in